
Cameo: A Python Library for Computer Aided Metabolic Engineering and Optimization of Cell Factories | ACS Synthetic Biology

Growth rate predictions for 22 single knockout E. coli mutants using... | Download Scientific Diagram

Machine learning-guided evaluation of extraction and simulation methods for cancer patient-specific metabolic models - ScienceDirect

Advances in flux balance analysis by integrating machine learning and mechanism-based models - ScienceDirect
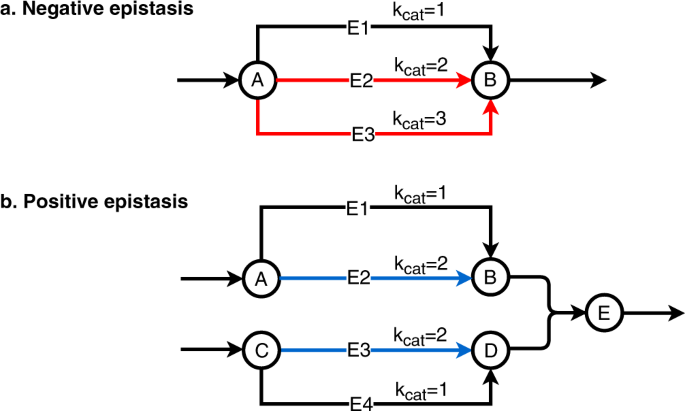
Flux balance analysis with or without molecular crowding fails to predict two thirds of experimentally observed epistasis in yeast | Scientific Reports

pFBA simulation of a single solution. Difference in simulation results... | Download Scientific Diagram

FBA, MOMA and PSEUDO approaches to predicting metabolic fluxes. (A) In... | Download Scientific Diagram

Conceptual architecture of the MEWpy framework. Phenotype prediction... | Download Scientific Diagram

Emergent sub-population behavior uncovered with a community dynamic metabolic model of Escherichia coli diauxic growth | bioRxiv
In silico co-factor balance estimation using constraint-based modelling informs metabolic engineering in Escherichia coli

The relationship between the target chemical production flux and the... | Download Scientific Diagram
ΔFBA—Predicting metabolic flux alterations using genome-scale metabolic models and differential transcriptomic data | PLOS Computational Biology

Plant genome-scale reconstruction: from single cell to multi-tissue modelling and omics analyses - ScienceDirect
Metabolic Adaptation Processes That Converge to Optimal Biomass Flux Distributions | PLOS Computational Biology

FBA, MOMA and PSEUDO approaches to predicting metabolic fluxes. (A) In... | Download Scientific Diagram

FBA, MOMA and PSEUDO approaches to predicting metabolic fluxes. (A) In... | Download Scientific Diagram
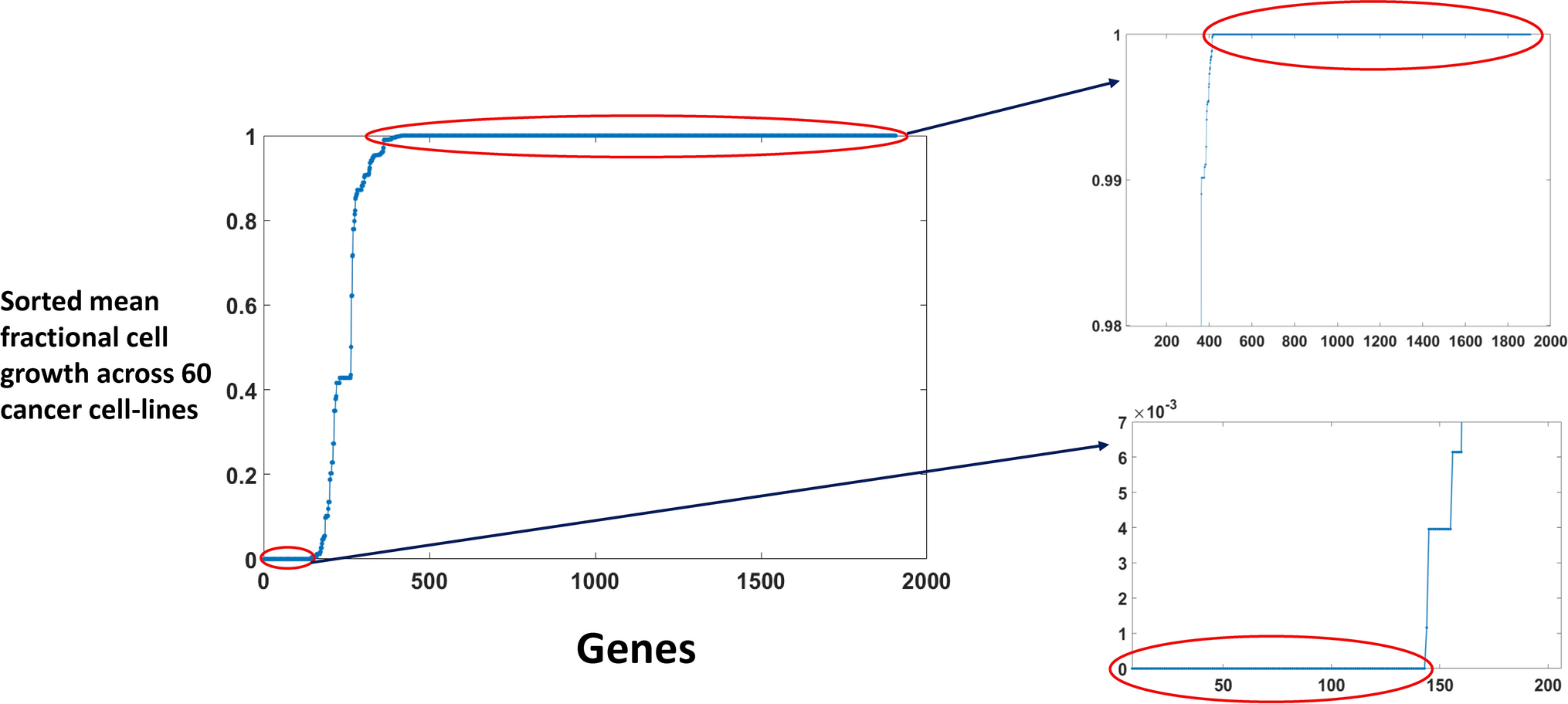
Exploring gene knockout strategies to identify potential drug targets using genome-scale metabolic models | Scientific Reports
Stoichiometric Representation of Gene–Protein–Reaction Associations Leverages Constraint-Based Analysis from Reaction to Gene-Level Phenotype Prediction | PLOS Computational Biology
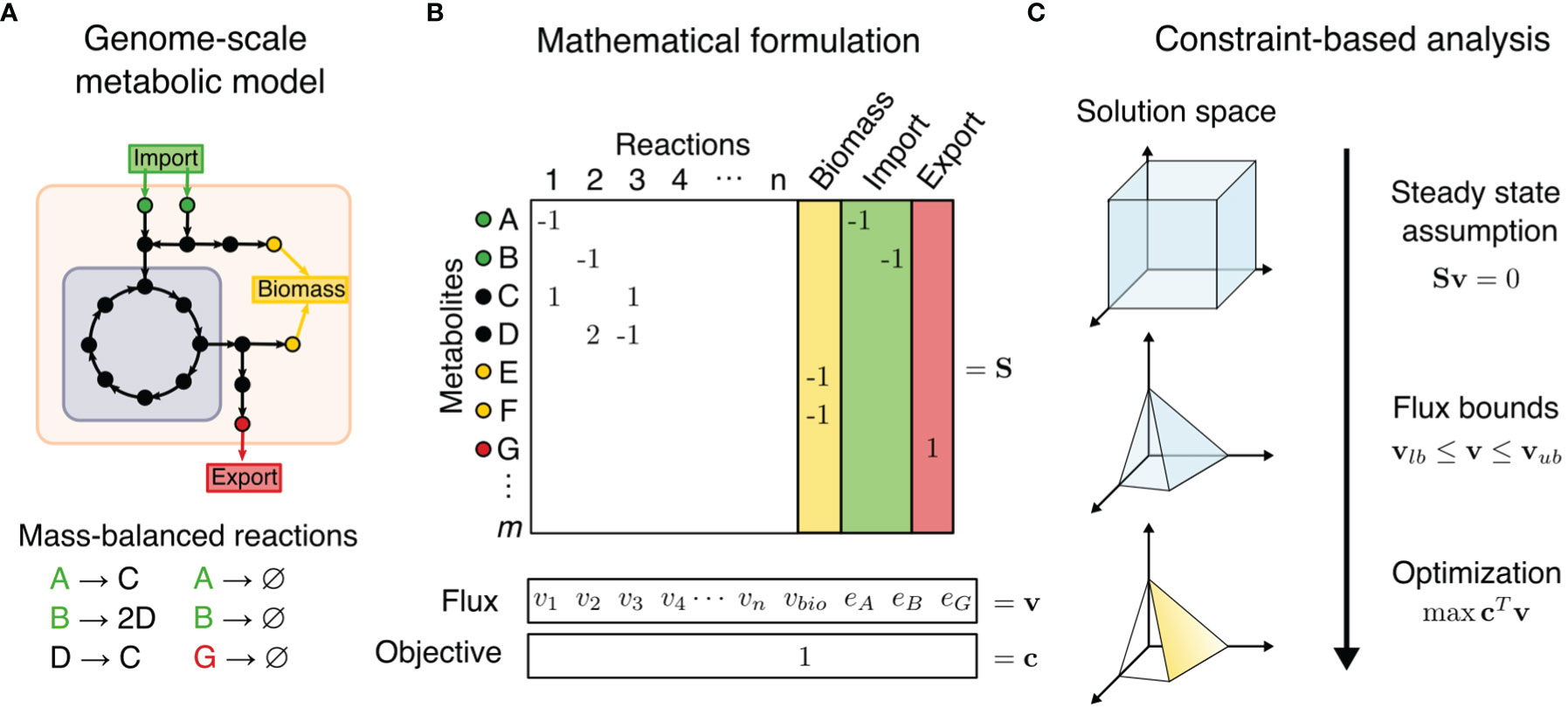
Frontiers | Constraint-Based Reconstruction and Analyses of Metabolic Models: Open-Source Python Tools and Applications to Cancer







